The human body is composed of billions of cells, each of which is made and maintained through countless interactions among its molecular parts. But which interactions sustain health and which ones can cause disease when they go awry? The human genome project has provided us with a “parts list” for the cell, but only if we can understand how these parts go together, or interact, can we really begin to understand how the cell works and what goes wrong in disease.
To answer these questions, scientists needed a reference map of interactions—an interactome— between gene-encoded proteins, which make up cells and do most of the work in them.
Almost a decade in the making, the human protein map is now available thanks to a joint effort, involving over 80 researchers in the United States, Canada, Spain, Belgium, France and Israel, led by Marc Vidal, David Hill and Michael Calderwood, at the Center for Cancer Systems Biology (CCSB) at Dana-Farber Cancer Institute, and Frederick Roth, a professor of molecular genetics and computer science at U of T’s Donnelly Centre.
The largest of its kind, the human reference interactome (HuRI) map charts 52,569 interactions between 8,275 human proteins, as described in a study published in Nature.
Read more at University of Toronto
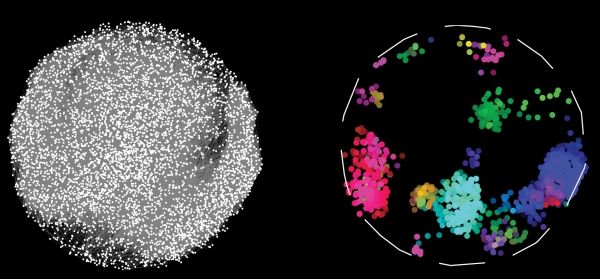
Image Credit: University of Toronto